

中国核心期刊要目总览
中国科技核心期刊
CSCD数据库源刊
中国科技核心期刊
CSCD数据库源刊


ACTA THERIOLOGICA SINICA ›› 2023, Vol. 43 ›› Issue (2): 182-192.DOI: 10.16829/j.slxb.150724
• ORIGINAL PAPERS • Previous Articles Next Articles
Weiwei SHAO, Fen QIAO, Wei CAI, Zhihua LIN, Li WEI( )
)
Received:2022-08-13
Accepted:2022-11-07
Online:2023-03-30
Published:2023-03-23
Contact:
Li WEI
通讯作者:
韦力
作者简介:邵伟伟 (1981- ),女,硕士,主要从事动物学研究.
基金资助:CLC Number:
Weiwei SHAO, Fen QIAO, Wei CAI, Zhihua LIN, Li WEI. Characteristics of microsatellite distributions in genomes of Hipposideros armiger (Chiroptera)[J]. ACTA THERIOLOGICA SINICA, 2023, 43(2): 182-192.
邵伟伟, 乔芬, 蔡玮, 林植华, 韦力. 大蹄蝠全基因组微卫星分布特征分析[J]. 兽类学报, 2023, 43(2): 182-192.
微卫星类型 Type of SSR | 微卫星数量 Number of SSRs | 长度 Length (bp) | 丰度Abundance (loci/Mb) | 比例 Percentage (%) |
|---|---|---|---|---|
| 单碱基 Mononucleotide | 173 953 | 2 451 423 | 77.78 | 34.94 |
| 二碱基 Dinucleotide | 222 591 | 6 114 126 | 99.52 | 44.71 |
| 三碱基 Trinucleotide | 37 566 | 800 058 | 16.8 | 7.55 |
| 四碱基 Tetranucleotide | 47 250 | 1 517 188 | 21.13 | 9.49 |
| 五碱基 Pentanucleotide | 13 134 | 310 285 | 5.87 | 2.64 |
| 六碱基 Hexanucleotide | 3 389 | 98 148 | 1.52 | 0.68 |
| SSR总数 Total of SSRs | 497 883 | 11 291 228 | 222.61 | 100 |
| 总基因组大小 Whole genome length (bp) | 2 236 581 172 | |||
| 微卫星含量 SSR content of genome (%) | 0.50 | |||
Table 1 Distribution of microsatellites in the genomes of Hipposideros armiger
微卫星类型 Type of SSR | 微卫星数量 Number of SSRs | 长度 Length (bp) | 丰度Abundance (loci/Mb) | 比例 Percentage (%) |
|---|---|---|---|---|
| 单碱基 Mononucleotide | 173 953 | 2 451 423 | 77.78 | 34.94 |
| 二碱基 Dinucleotide | 222 591 | 6 114 126 | 99.52 | 44.71 |
| 三碱基 Trinucleotide | 37 566 | 800 058 | 16.8 | 7.55 |
| 四碱基 Tetranucleotide | 47 250 | 1 517 188 | 21.13 | 9.49 |
| 五碱基 Pentanucleotide | 13 134 | 310 285 | 5.87 | 2.64 |
| 六碱基 Hexanucleotide | 3 389 | 98 148 | 1.52 | 0.68 |
| SSR总数 Total of SSRs | 497 883 | 11 291 228 | 222.61 | 100 |
| 总基因组大小 Whole genome length (bp) | 2 236 581 172 | |||
| 微卫星含量 SSR content of genome (%) | 0.50 | |||
模体单元 Motif length | 重复单元 Repeat unit | 微卫星数量 Microsatellites | 比例 Percentage (%) |
|---|---|---|---|
单碱基重复 Mononucleotide repeat | A | 165 494 | 95.14 |
| G | 8 459 | 4.86 | |
二碱基重复 Dinucleotide repeat | AC | 122 977 | 55.25 |
| CT | 42 659 | 19.16 | |
| GC | 1 460 | 0.66 | |
| TA | 55 495 | 24.93 | |
三碱基重复 Trinucleotide repeat | TAT | 14 429 | 38.41 |
| CAA | 7 164 | 19.07 | |
| ACC | 3 653 | 9.72 | |
| CAT | 3 408 | 9.07 | |
四碱基重复 Tetranucleotide repeat | TTTA | 10 475 | 22.17 |
| AAAC | 8 049 | 17.03 | |
| ATAG | 7 078 | 14.98 | |
| CATT | 4 959 | 10.50 | |
五碱基重复 Pentanucleotide repeat | AACAA | 6 393 | 48.68 |
| TTATT | 2 544 | 19.37 | |
| TTTCT | 795 | 6.05 | |
| TATTA | 354 | 2.70 | |
六碱基重复 Hexanucleotide repeat | TATCTA | 688 | 20.30 |
| AAACAA | 510 | 15.05 | |
| GAGAGG | 284 | 8.38 | |
| AATAAA | 133 | 3.92 |
Table 2 The most frequent microsatellite motifs found in genomes of Hipposideros armiger
模体单元 Motif length | 重复单元 Repeat unit | 微卫星数量 Microsatellites | 比例 Percentage (%) |
|---|---|---|---|
单碱基重复 Mononucleotide repeat | A | 165 494 | 95.14 |
| G | 8 459 | 4.86 | |
二碱基重复 Dinucleotide repeat | AC | 122 977 | 55.25 |
| CT | 42 659 | 19.16 | |
| GC | 1 460 | 0.66 | |
| TA | 55 495 | 24.93 | |
三碱基重复 Trinucleotide repeat | TAT | 14 429 | 38.41 |
| CAA | 7 164 | 19.07 | |
| ACC | 3 653 | 9.72 | |
| CAT | 3 408 | 9.07 | |
四碱基重复 Tetranucleotide repeat | TTTA | 10 475 | 22.17 |
| AAAC | 8 049 | 17.03 | |
| ATAG | 7 078 | 14.98 | |
| CATT | 4 959 | 10.50 | |
五碱基重复 Pentanucleotide repeat | AACAA | 6 393 | 48.68 |
| TTATT | 2 544 | 19.37 | |
| TTTCT | 795 | 6.05 | |
| TATTA | 354 | 2.70 | |
六碱基重复 Hexanucleotide repeat | TATCTA | 688 | 20.30 |
| AAACAA | 510 | 15.05 | |
| GAGAGG | 284 | 8.38 | |
| AATAAA | 133 | 3.92 |
指标 Index | 基因区Genetic region | 基因间区 Intergenic region | |||
|---|---|---|---|---|---|
编码区 CDS | 非翻译区 Untranslated | 外显子 Exon | 内含子 Intron | ||
数量 Number | 1 461 | 5 081 | 7 416 | 199 340 | 322 666 |
丰度 Abundance(loci/Mb) | 461.98 | 1 647.13 | 1 188.18 | 2 204.77 | 2 541.57 |
Table 3 The number and abundance of microsatellites in different genomic regions of Hipposideros armiger
指标 Index | 基因区Genetic region | 基因间区 Intergenic region | |||
|---|---|---|---|---|---|
编码区 CDS | 非翻译区 Untranslated | 外显子 Exon | 内含子 Intron | ||
数量 Number | 1 461 | 5 081 | 7 416 | 199 340 | 322 666 |
丰度 Abundance(loci/Mb) | 461.98 | 1 647.13 | 1 188.18 | 2 204.77 | 2 541.57 |
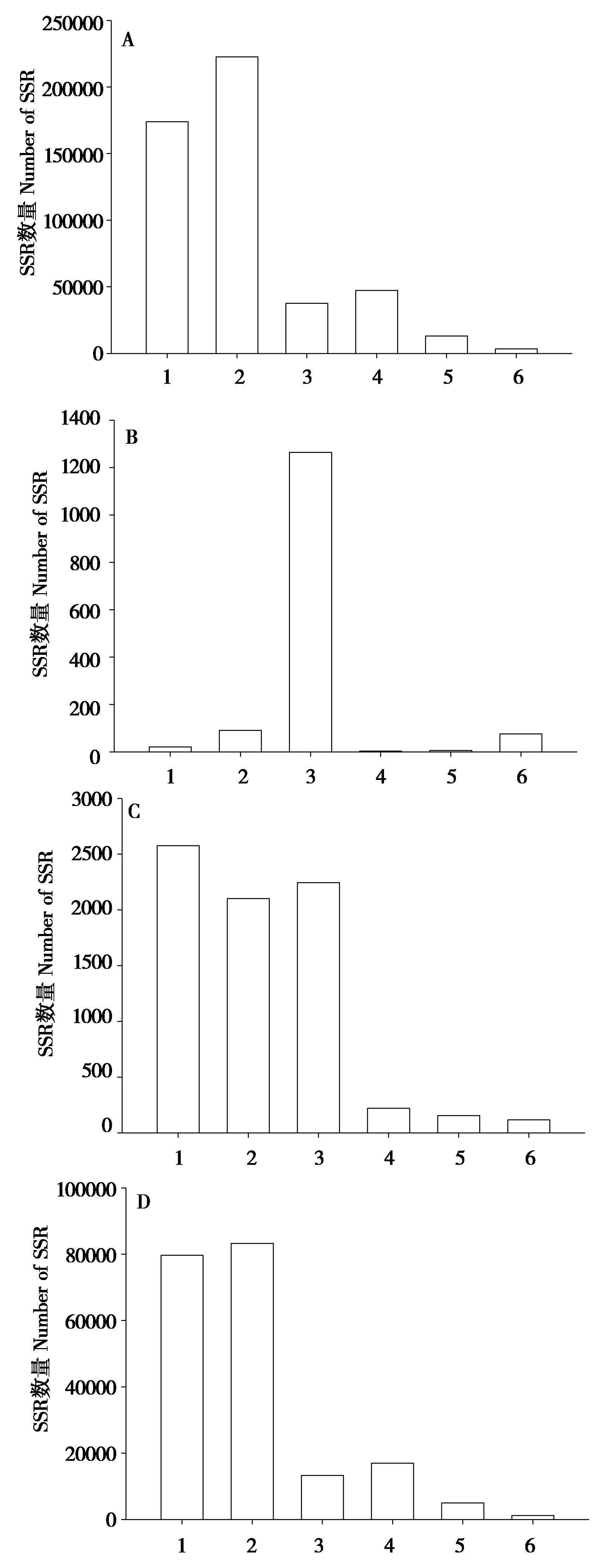
Fig. 2 The distribution of microsatellite types in different genomic regions.Genome (A), CDS (B), Exon (C) and Intergenetic (D) of Hipposideros armiger. 1 - 6 indicated mono-, di-, tri-, tetra-, penta-, hexa-nucleotide respectively
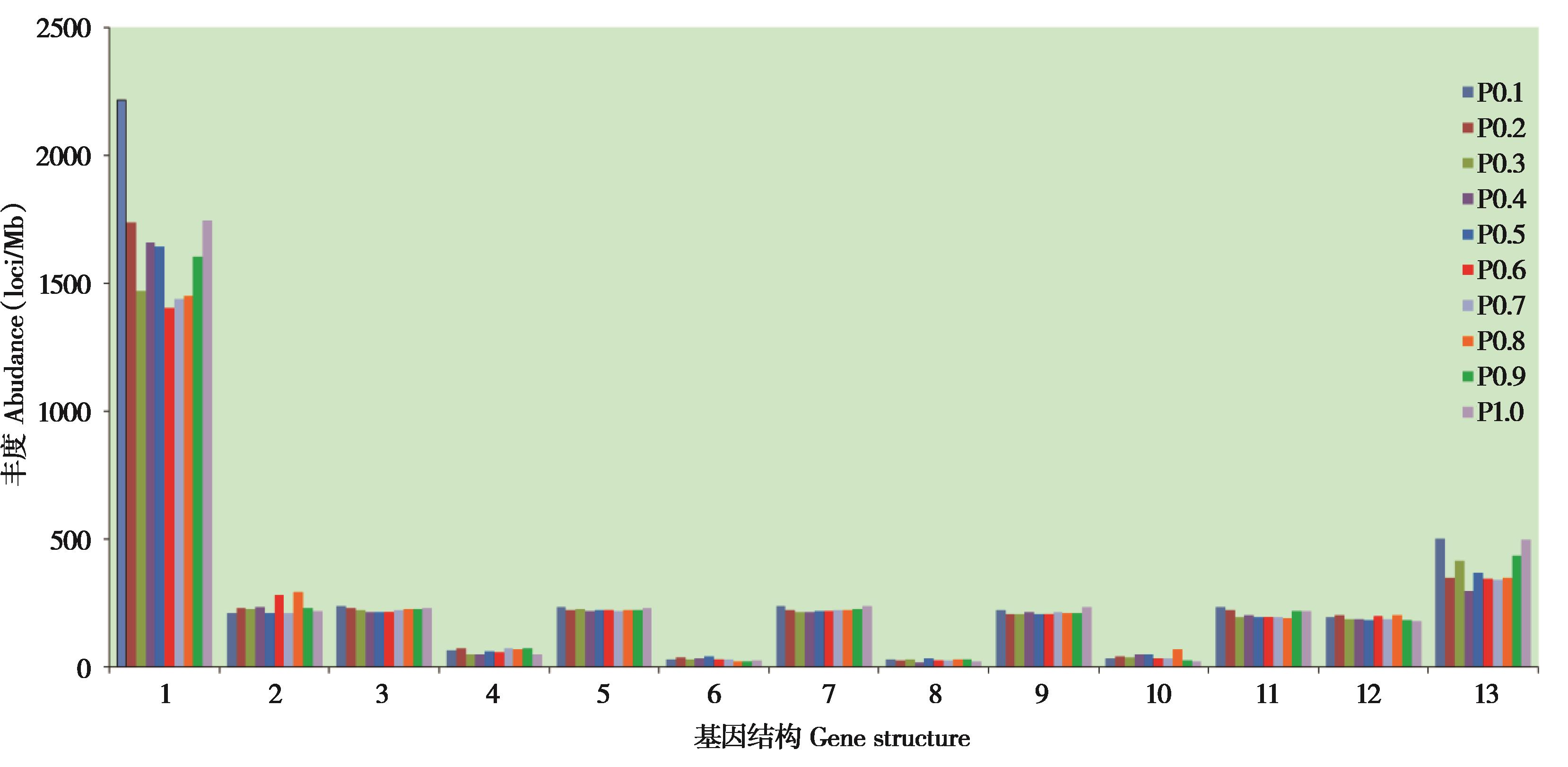
Fig. 3 The microsatellite abundance in gene regions and their upstream and downstream regions of Hipposideros armiger. Number 1 - 13 indicated upstream 500 bp, exon 1, intron1, exon 2, intron 2, mid left exon, mid intron, mid right exon,intron reverse 2, exon reverse 2, intron reverse 1, exon reverse 1, downstream 500 bp, respectively
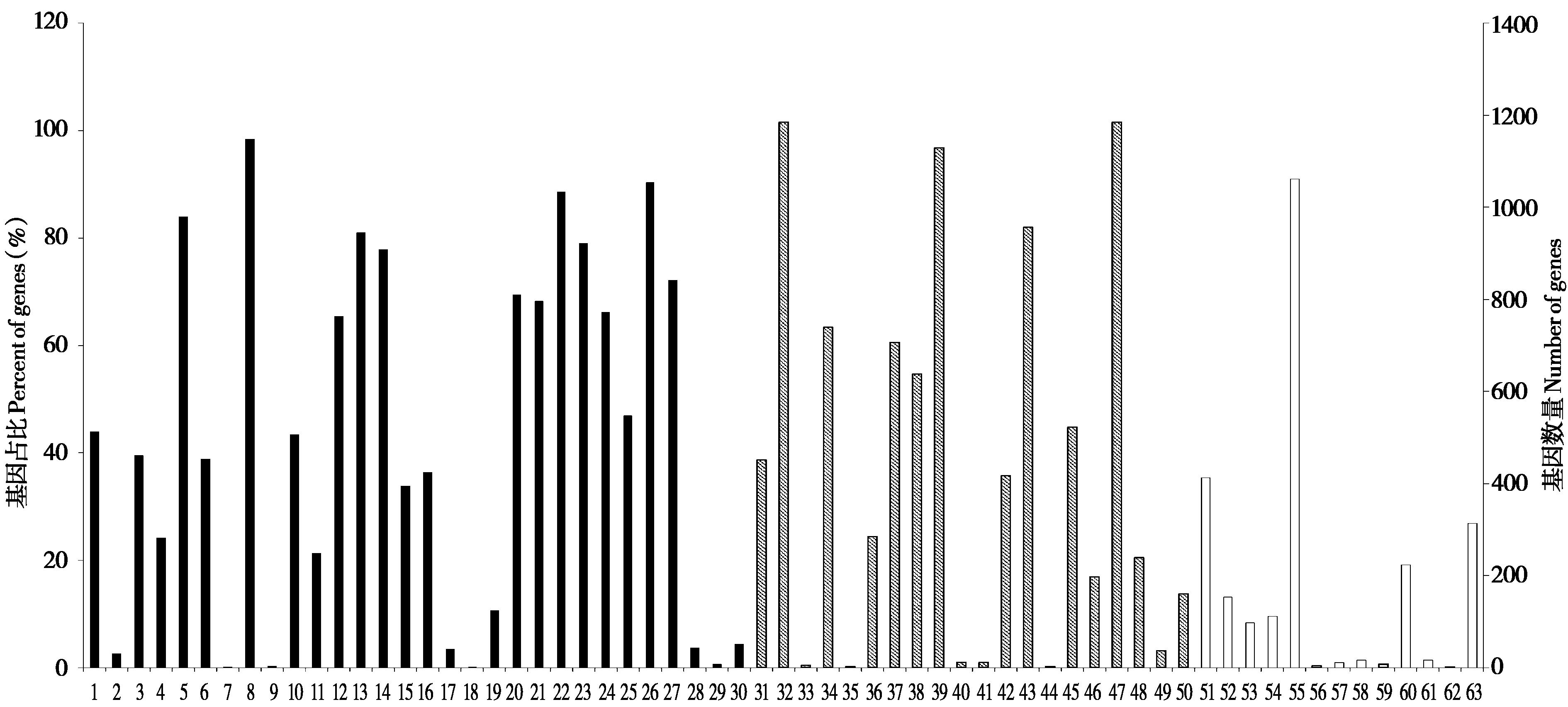
Fig. 4 GO classifications of coding sequencings with microsatellites in the genomes of Hipposideros armiger. 1: Reproduction; 2: Cell killing; 3: Immune system process; 4: Behavior; 5: Metabolic process; 6: Cell proliferation; 7: Carbohydrate utilization; 8: Cellular process; 9: Nitrogen utilization; 10: Reproductive process; 11: Biological adhesion; 12: Signaling; 13: Multicellular organismal process; 14: Developmental process; 15: Growth; 16: Locomotion; 17: Pigmentation; 18: Biological phase; 19: Rhythmic process; 20: Positive regulation of biological process; 21: Negative regulation of biological process; 22: Regulation of biological process; 23: Response to stimulus; 24: Localization; 25: Multi-organism process; 26: Biological regulation; 27: Cellular component organization or biogenesis; 28: Cell aggregation; 29: Detoxification; 30: Presynaptic process involved in chemical synaptic transmission; 31: Extracellular region; 32: Cell; 33: Nucleoid; 34: Membrane; 35: Virion; 36: Cell junction; 37: Membrane-enclosed lumen; 38: Protein-containing complex; 39: Organelle; 40: Other organism; 41: Other organism part; 42: Extracellular region part; 43: Organelle part; 44: Virion part; 45: Membrane part; 46: Synapse part; 47: Cell part; 48: Synapse; 49: Symplast; 50: Supramolecular complex; 51: Catalytic activity; 52: Signal transducer activity; 53: Structural molecule activity; 54: Transporter activity; 55: Binding; 56: Antioxidant activity; 57: Cargo receptor activity; 58: Translation regulator activity; 59: Toxin activity; 60: Molecular function regulator; 61: Hijacked molecular function; 62: Molecular carrier activity; 63: Transcription regulator activity. The black, stripe and white area indicated biological process, cellular component and molecular function, respectively
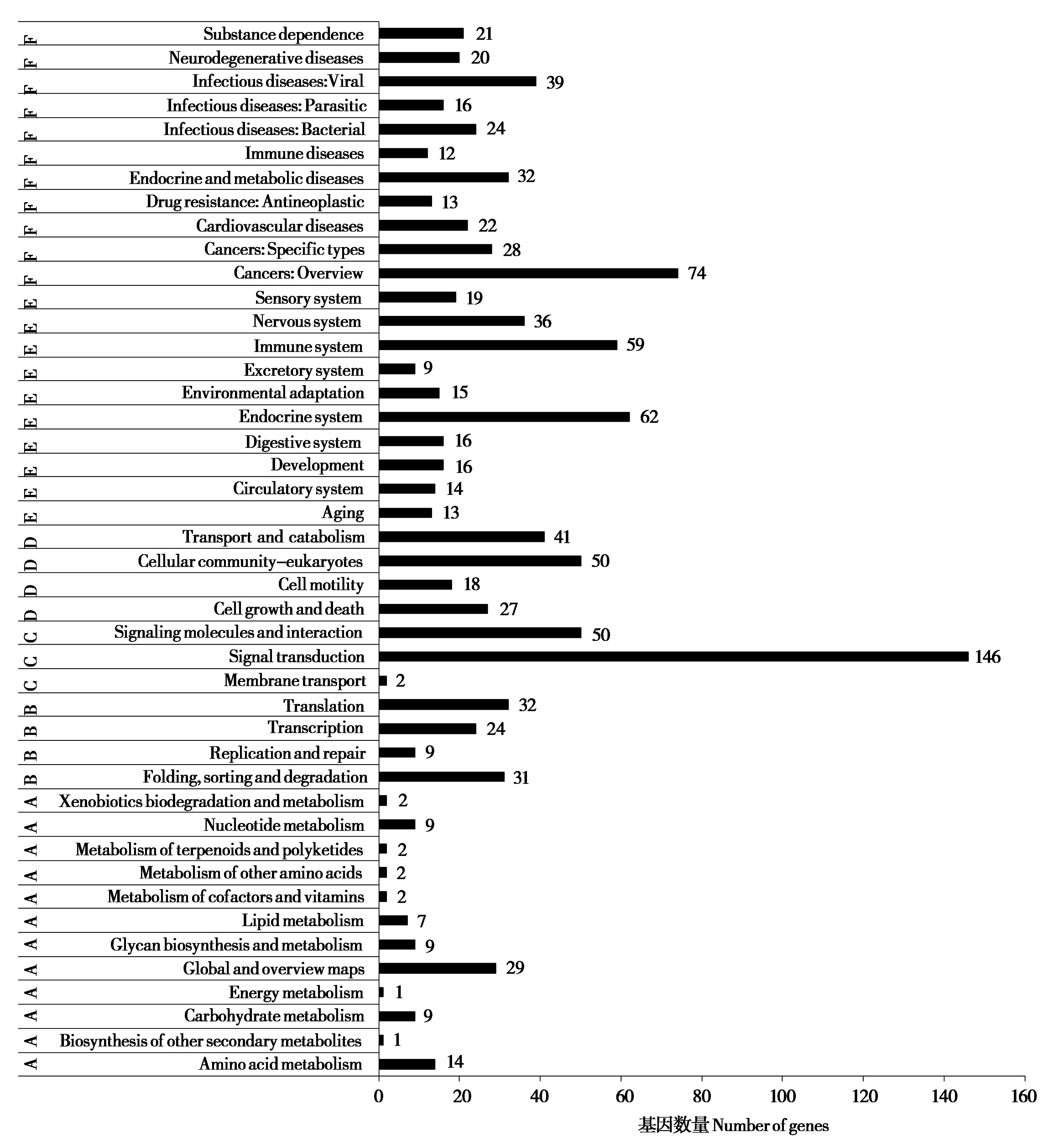
Fig. 5 The KEGG enrichment of exon microsatellites in Hipposideros armiger. A: Metabolism; B: Environmental information processing; C: Genetic information processing; D: Cell process; E: Organismal systems; F: Human diseases and drug development
| Castillo‑Lizardo M, Henneke G, Viguera E. 2014. Replication slippage of the thermophilic DNA polymerases B and D from the Euryarchaeota Pyrococcus abyssi . Frontiers in Microbiology, 5 (403): 403-403. | |
| Collaborative R. 2022. Impact of microsatellite status in early-onset colonic cancer. British Journal of Surgery, 109 (7): 632-636. | |
| Cui K, Yue B S. 2018. Distribution patterns of microsatellites in the genome of Lophophorus lhuysii . Sichuan Journal of Zoology, 37 (5): 533-540. (in Chinese) | |
| Deback C, Boutolleau D, Depienne C, Luyt C E, Bonnafous P, Gautheret‑Dejean A, Garrigue I, Agut H. 2009. Utilization of microsatellite polymorphism for differentiating herpes simplex virus type 1 strains. Journal of Clinical Microbiology, 47: 533-540. | |
| Dhyani P, Sharma B, Singh P, Masand M, Sharma R. 2020. Genome-wide discovery of microsatellite markers and, population genetic diversity inferences revealed high anthropogenic pressure on endemic populations of Trillium govanianum . Industrial Crops and Products, 154: 112698. | |
| Ding S M, Wang S P, He K, Jiang M X, Li F. 2017. Large scale analysis reveals that the genome features of simple sequence repeats are generally conserved at the family level in insects. BMC Genomics, 18: 848. | |
| Ellegren H. 2004. Microsatellites: simple sequences with complex evolution. Nature Reviews Genetics, 5 (6): 435-445. | |
| Fan S G, Huang H, Liu Y, Wang P F, Zhao C, Yan L L, Qiao X T, Qiu L H. 2021. Genome‑wide identification of microsatellite and development of polymorphic SSR markers for spotted sea bass (Lateolabrax maculatus). Aquaculture Reports, 20: 100677. | |
| Fujimori S, Washio T, Higo K. 2003. A novel feature of microsatellites in plants: a distribution gradient among the direction of transcription. FEBS Letters, 554 (1): 17-22. | |
| Gao H, Liu P, Meng X H, Wang W J, Kong J. 2004. Analysis of microsatellite sequences in Chinese shrimp Fenneropenaeus chinensis genome. Oceanologia et Limnologia Sinica, 35 (5): 424-431. (in Chinese) | |
| George B, Gnanasekaran P, Jain S K, Chakraborty S. 2014. Genome wide survey and analysis of small repetitive sequences in caulimoviruses. Infection Genetics & Evolution, 27: 15-24. | |
| Hancock J M. 1996. Simple sequences in a ‘minimal’ genome. Nature Genetics, 14: 14-15. | |
| Huang J, Du L M, Li Y Z, Li W J, Zhang X Y, Yue B S. 2012. Distribution regularities of microsatellites in the Gallus gallus genome. Sichuan Journal of Zoology, 31 (3): 358-363. (in Chinese) | |
| Huntley M A, Golding G B. 2006. Selection and slippage creating serine homopolymers. Molecular Biology and Evolution, 23: 2017-2025. | |
| Kashi Y, King D G. 2006. Simple sequence repeats as advantageous mutators in evolution. Trends in Genetics, 22: 253-259. | |
| Li W J, Li Y Z, Du L M, Huang J, Shen Y M, Zhang X Y, Yue B S. 2014. Comparative analysis of microsatellite sequences distribution in the genome of giant panda and polar bear. Sichuan Journal of Zoology, 33 (6): 874-878. (in Chinese) | |
| Lin W H, Kussel E. 2012. Evolutionary pressures on simple sequence repeats in prokaryotic coding regions. Nucleic Acids Research, 40: 2399-2413. | |
| Loire E, Higuet D, Netter P, Achaz G. 2013. Evolution of coding microsatellites in primate genomes. Genome Biology and Evolution, 5: 283-295. | |
| Lu K M, Wang T J, Dong S W, Xing X M, Su W L. 2022. Research on distribution characteristics of microsatellites in Sika deer genome. Special Wild Economic Animal and Plant Research, 44 (6): 8-15, 23.(in Chinese) | |
| Lu T, Wang C, Du C, Liu S, Shen Y M, Zhang X Y, Yue B S. 2017. Distribution regularity of microsatellites in Moschus berezovskii genome. Sichuan Journal of Zoology, 36 (4): 420-424. (in Chinese) | |
| Mayer C, Leese F, Tollrian R. 2010. Genome‑wide analysis of tandem repeats in Daphnia pulex–a comparative approach. BMC Genomics, 11: 277. | |
| Miah G, Rafii M Y, Ismail M R, Puteh A B, Rahim H A, Islam K N, Latif M A. 2013. A Review of microsatellite markers and their applications in rice breeding programs to improve blast disease resistance. International Journal of Molecular Science, 14: 22499-22528. | |
| Mirkin S M. 2007. Expandable DNA repeats and human disease. Nature, 447: 932-940. | |
| Nie H, Cao S S, Zhao M L, Du L F. 2017. Comparative analysis of microsatellite distributions in genomes of Boa constrictor and Protobothrops mucrosquamatus . Sichuan Journal of Zoology, 36 (6): 639-648.(in Chinese) | |
| Raghunath S. 2022. Application of Bioinformatics resources for mining of simple sequence repeats (SSRs) marker in plant genomes: an overview. Research Journal of Biotechnology, 17 (8): 136-143. | |
| Rogers J, Gibbs R A. 2014. Comparative primate genomics: emerging patterns of genome content and dynamics. Nature Reviews Genetics, 15: 347-359. | |
| Smith T H, Xie Y. 2008. A Guide to the Mammals of China. Princeton, NJ: Princeton University Press. | |
| Subramanian S, Mishra R K, Singh L. 2003. Genome‑wide analysis of microsatellite repeats in humans: their abundance and density in specific genomic regions. Genome Biology, 4 (2): 1-10. | |
| Tay W T, Behere G T, Batterham P, Heckel D G. 2010. Generation of microsatellite repeat families by RTE retrotransposons in lepidopteran genomes. BMC Evolutionary Biology, 10: 144. | |
| Tóth G, Góspóri Z, Jurka J. 2000. Microsatellites in different eukaryotic genomes: survey and analysis. Genome Research, 10: 967-981. | |
| Tu F Y, Liu J, Han W J, Huang T, Huang X F. 2018. Analysis of microsatellite distribution characteristics in the entire genome of Macaca fascicularis . Chinese Journal of Wildlife, 39 (2): 400-404. (in Chinese) | |
| Walter R, Epperson B K. 2001. Geographic pattern of genetic variation in Pinus resinosa: area of greatest diversity is not the origin of postglacial populations. Molecular Ecology, 10: 103-111. | |
| Wang J, Luo Q, Duan Q, Zhong K L, Huang X T, Zhang R Y. 2019. Screening and analysis of microsatellites in the genomes of three Sinocyclocheilus fishes. Journal of Guizhou Normal University (Natural Sciences), 37 (4): 19-24. (in Chinese) | |
| Wang Y Y, Liu X X, Dong K Z, Chen X F, Ye S H, Ma Y H. 2015. Distribution difference of microsatellite in 7 domestic animals genome. China Animal Husbandry & Veterinary Medicine, 42 (9): 2418-2426. (in Chinese) | |
| Wei L, Shao W W, Ma L, Lin Z H. 2020.Genome wide analysis of microsatellite markers based on sequenced database in two anuran species. Journal of Genetics, 99: 58. | |
| Yin W, Wang Z J, Li Q Y, Lian J M, Zhou Y, Lu B Z, Jin L J, Qiu P X, Zhang P, Zhu W B, Wen B, Huang Y J, Lin Z L, Qiu B T, Su X W, Yang H M, Zhang G J, Yan G M, Zhou Q.2016. Evolutionary trajectories of snake genes and genomes revealed by comparative analysis of five‑pacer viper. Nature Communications, 13107 (1):1-11. | |
| Zane L, Bargelloni L, Patarnello T. 2002. Strategies for microsatellite isolation: a review. Molecular Ecology, 11: 1-16. | |
| 王月月, 刘雪雪, 董坤哲, 陈潇飞, 叶绍辉, 马月辉. 2015. 7种家养动物全基因组微卫星分布的差异研究. 中国畜牧兽医, 42 (9): 2418-2426. | |
| 王军, 罗琦, 段茜, 钟凯丽, 黄晓彤, 张仁意. 2019. 3种金线鲃属鱼类基因组微卫星的筛选与分析. 贵州师范大学学报 (自然科学版), 37 (4): 19-24. | |
| 卢凯妹, 王天骄, 董世武, 邢秀梅, 苏伟林. 2022. 梅花鹿基因组微卫星分布特征研究. 特产研究, 44 (6): 8-15, 23. | |
| 卢婷, 王晨, 杜超, 刘姝, 沈咏梅, 张修月, 岳碧松. 2017. 林麝全基因组微卫星分布规律研究.四川动物, 36 (4): 420-424. | |
| 李午佼, 李玉芝, 杜联明, 黄杰, 沈咏梅, 张修月, 岳碧松. 2014. 大熊猫和北极熊基因组微卫星分布特征比较分析. 四川动物, 33 (6): 874-878. | |
| 聂虎, 曹莎莎, 赵明朗, 杜林方. 2017.红尾蚺和原矛头蝮基因组微卫星分布特征比较分析.四川动物, 36 (6): 639-648. | |
| 高焕, 刘萍, 孟宪红, 王伟继, 孔杰. 2004. 中国对虾 (Fenneropenaeus chinensis) 基因组微卫星特征分析. 海洋与湖沼, 35 (5): 424-431. | |
| 涂飞云, 刘俊, 韩卫杰, 黄挺, 黄晓凤. 2018. 食蟹猴全基因组微卫星分布特征分析. 野生动物学报, 39 (2): 400-404. | |
| 黄杰, 杜联明, 李玉芝, 李午佼, 张修月, 岳碧松. 2012. 红原鸡全基因组中微卫星分布规律研究. 四川动物, 31 (3): 358-363. | |
| 崔凯, 岳碧松. 2018. 绿尾虹雉全基因组微卫星分布规律研究. 四川动物, 37 (5): 533-540. |
| [1] | Lü Zelong, LI Kexin, HU Yibo. Research on genetic diversity and global distribution pattern of feline species [J]. ACTA THERIOLOGICA SINICA, 2024, 44(6): 795-803. |
| [2] | ZHANG Yang, CHEN Luyao, HAN Weijie, ZHAN Jianwen, LIU Wuhua, HUANG Xiaofeng. Genetic diversity of sika deer based on microsatellite in Taohongling, Jiangxi [J]. ACTA THERIOLOGICA SINICA, 2024, 44(3): 268-276. |
| [3] | ZHENG Kaidan, WANG Qiaoyun, FAN Pengfei, HAN Xuesong, XIAO Mei, SHEN Limin, DONG Zhengyi, ZHANG Lu. Individual identification and genetic diversity of Eurasian otters based on microsatellite markers [J]. ACTA THERIOLOGICA SINICA, 2024, 44(2): 146-158. |
| [4] | HUANG Zefeng, LIAO Yaqing, WANG Xiaoyun, ZHANG Huiguang, CAI Bin, YONG Fan, CUI Peng, YU Wenhua, WU Yi. New records of two tube-nosed bats (CHIROPTERA: Vespertilionidae: Murina) from Fujian Province, China [J]. ACTA THERIOLOGICA SINICA, 2023, 43(4): 472-478. |
| [5] | TIAN Xinmin, ZHANG Minghai. Genetic diversity of wapiti in northeast China based on fecal DNA [J]. ACTA THERIOLOGICA SINICA, 2023, 43(1): 41-49. |
| [6] | LI Yannan, LIANG Xiaoling, XIE Huixian, DENG Wenpu, HE Minyi, YU Wenhua, WU Yi. Rediscovery of Theobald's tomb bat (Taphozous theobaldi, Chiroptera, Emballonuridae) from Guangdong, China [J]. ACTA THERIOLOGICA SINICA, 2023, 43(1): 122-128. |
| [7] | Xinmin TIAN, Mingdong LIAN, Yaqi SONG, Xiaohui LIU, Mengping YANG, Hong CHEN. Genetic diversity and demographic history of Siberian flying squirrel (Pteromys volans) population in northern Zhangguangcai Mountains, Heilongjiang, China [J]. ACTA THERIOLOGICA SINICA, 2022, 42(4): 398-409. |
| [8] | WANG Junhua, WONG Kai-Chin, CHEK Si-Nga, VU Ka-Man, CHAN Hoi-Hou, LIANG Jie, HE Xiangyang, ZHANG Libiao. New records of bat species and their conservation status in Macao, China [J]. ACTA THERIOLOGICA SINICA, 2022, 42(1): 125-130. |
| [9] | XIE Huixian, LI Yannan, LIANG Xiaoling, ZHANG Huiguang, ZHAN Liying, WU Yi, YU Wenhua. Second record of collared sprite(Thainycteris aureocollaris, Chiroptera, Vespertilionidae) from China [J]. ACTA THERIOLOGICA SINICA, 2021, 41(4): 476-482. |
| [10] | ZHANG Zhao, ZHANG Rui, LI Xiaoyu, Saihan, YANG Zhendong, HAN Zhiqing, BAO Weidong. Genetic diversity and sex structure of red deer population in Saihanwula Nature Reserve, Inner Mongolia [J]. ACTA Theriologica Sinica, 2021, 41(1): 42-50. |
| [11] | WANG Dou, XU Guan, WANG Hongyong, HE Sen, BU Shuhai, ZHENG Xueli. Study on polymorphisms of microsatellites DNA of Chinese captive forest musk deer(Moschus berezovskii) [J]. ACTA THERIOLOGICA SINICA, 2019, 39(6): 599-607. |
| [12] | ZHANG Yuguang, Charlotte HACKER, ZHANG Yu, XUE Yadong, WU Liji, DAI Yunchuan, LUO Ping, Xierannima, Jan E JANECKA, LI Diqiang. An analysis of genetic structure of snow leopard populations in Sanjiangyuan and Qilianshan National Parks [J]. ACTA THERIOLOGICA SINICA, 2019, 39(4): 442-449. |
| [13] | PANG Yulan, LUO Bo, WANG Man, WU Xiu, FENG Jiang. Sexual dimorphism in the frequency of echolocation calls facilitates sex recognition in least horseshoe bats [J]. ACTA THERIOLOGICA SINICA, 2019, 39(2): 155-161. |
| [14] | QIAO Maiju, RAN Jianghong, ZHANG Hemin . The application of microsatellite markers in giant panda research [J]. ACTA THERIOLOGICA SINICA, 2019, 39(1): 103-110. |
| [15] | XIU Yunfang, LIU Guowei, ZHENG Shuhuan. Genetic diversity and population genetic structure of captive red pandas (Ailurus fulgens) [J]. ACTA THERIOLOGICA SINICA, 2018, 38(4): 393-401. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
 青公网安备 63010402000199号
青ICP备05000010号-2
青公网安备 63010402000199号
青ICP备05000010号-2